#pedigree
Text
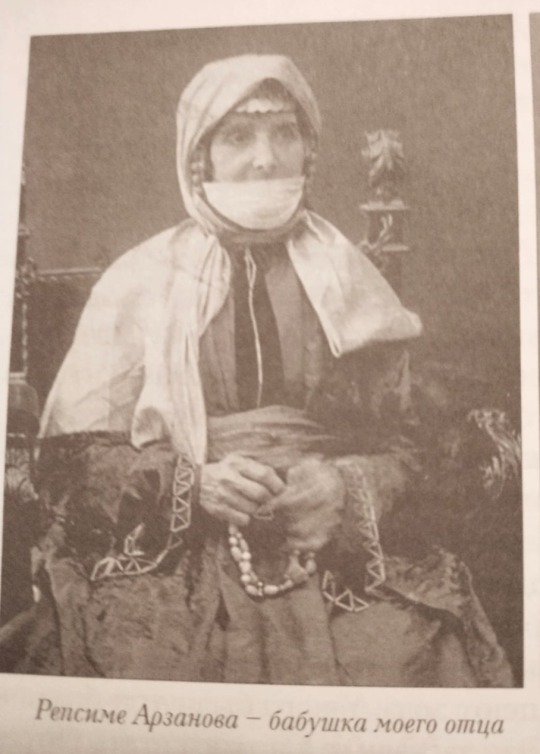
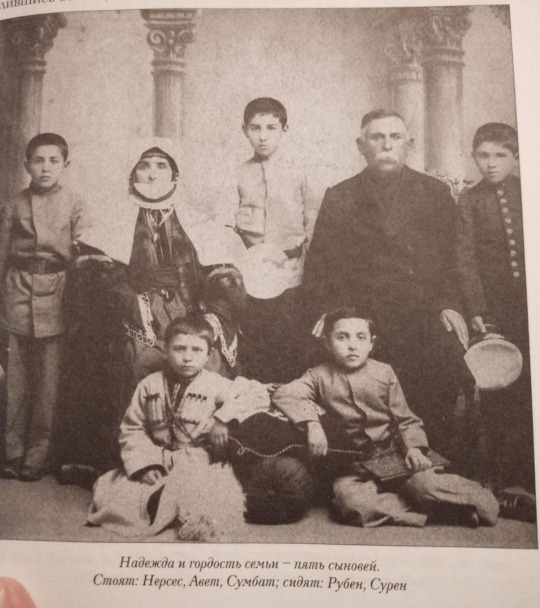


a photograph of my great-great-great grandmother, Rapsime Arzanova, and her husband with their 5 sons. those that stand: Nerses, Avet and Sumbat (my great-grandfather). Those who are sitting: Ruben and Suren.
#art#assets#русский художник#illustration#family history#history#caucasus#armenians#armenia#Armenian family#old photos#pedigree#Caucasian culture#Armenian culture#nation#nationality#National clothes#folk costume#folk clothes
33 notes
·
View notes
Text







I care a lot, 2020
7 notes
·
View notes
Text

I took this photo as part of the theme #thinkpink for #mamsellemagazine as part of #sindyat60 I hope that you like it. Image Description: a photo of a vintage Sindy dressed in a pink party dress sitting in her living room. There is a slice of #birthdaycake on a plate to her side and there is a table on it with two glasses one pink and one with champagne in it, a bucket of ice with a champagne bottle in it and a copy of Mam’selle on it. #vintagesindydolls #vintagesindy #vintagesindydolls #pedigreesindy #sindy #dollphotography #party #miniatures #pink #dolls #vintagesindydoll #dresslikeasindy
#dolls#sindy#sindy doll#sindy dolls#pedigree#pedigree sindy#vintage sindy#vintage dolls#doll photography#my doll photos#miniatures#party#Sindy at 60#dollblr#doll collecting#doll collector
13 notes
·
View notes
Text
ROMAN SPEARED THE ROCK OMEBSYWHENEO
4 notes
·
View notes
Text
Let's Talk About Degrees — the Pedigree!
Hello Shih Tzu buddies!
Time for a heart-to-heart about something crucial – what keeps our little buddies wagging their tails? We're diving into the world of Dog Food, in this case the famous Pedigree brand. And trust us, it's not just any dog chow – it's a game-changer, especially for our Shih Tzus.

Pedigree has been a reliable name in the dog food game, and it's earned that reputation for a reason. Dedicated Pedigree customers have said that, even after experimenting with other brands, their dogs in particular choose Pedigree more often their own. With a history steeped in providing top-notch dog nutrition, they have a keen understanding of what our furry friends need. What makes Pedigree stand out is its dedication to tailoring nutrition for different breeds, making it an excellent option for our Shih Tzus.
Their Small Dog Complete Nutrition formula, in particular, caught our eye. Its’ formula boasts a blend of antioxidants, vitamins, and minerals that support overall health. And let's not forget the Omega-6 fatty acids for a shiny coat and a special fiber blend for healthy digestion – two things our Shih Tzus absolutely adore!

For those who love digging into the nitty-gritty details, Pedigree breaks it down on their website [https://www.pedigree.com/dog-care-articles/6-ways-help-your-small-dog-live-large]
Of course, different individuals have different experiences, and it's no different when it comes to pet food brands like Pedigree. We've come across diverse feedback, and it's important to acknowledge that not everyone might share the same perspective on Pedigree. Choosing the right pet food involves a bit of trial and error, as well as paying attention to your pet's individual response to different brands and formulations. Remember, as responsible pet owners, it's crucial to recognize that what works wonders for one pet may not necessarily be the perfect fit for another.
2 notes
·
View notes
Text

Pedigree Vol.2
8 notes
·
View notes
Quote
Say not that you have royal blood in thy veins, and are born of God, except you can prove your pedigree by daring to be holy.
William Gurnall
#holiness (lacking)#false profession#pedigree#royal blood#child of God#holy living#William Gurnall#Christian Quotes
9 notes
·
View notes
Text

#today's words were#yelp#vampire#pedigree#the case study of vanitas#vanitas no carte#vnc#les memoires de vanitas#noe archiviste#vnc murr
6 notes
·
View notes
Video
Sky blue by dollyfan1
43 notes
·
View notes
Text
I took a Pedigree! 😱😱😱
#pro wrestler#trans woman#transgender#wrestler#wrestling#aew#trans#trans wrestler#woman wrestler#wwe#hhh#triple h#pedigree#finisher
3 notes
·
View notes
Video
youtube
Biscrok Pedigree é boa? Biscoito Pedigree Biscrok Para Cães Adultos Mult...
4 notes
·
View notes
Text

I’m sorry for my absence I returned home this week after a fantastic holiday in Spain with my parents. I took this photo of Rachel my #customsindy doll who was customised by dollybirdsdolls with me and took this photo of her whilst in Spain. I feel incredibly fortunate to have gone on holiday. I hope that you like this photo. Image Description: a photo of a #sindydoll dressed in #summerclothes , holding a suitcase and standing on a green and white ceramic tile bench. #spain #sindy #sindydolls #customsindydoll #ooakdoll #sindycollector #holiday #dollphotography #dolls #pedigreesindy #spaintravel #dollcollector #dollcollecting #sindycollection
#dolls#Rachel#sindy doll#Sindy#Sindy dolls#Spain#holiday#pedigree#pedigree sindy#doll photography#dollblr#vintage doll#ooak#ooak sindy#ooak doll#vintage#my doll photos#doll collector#doll collection#doll collecting
5 notes
·
View notes
Text
Can a 6 cM connection be meaningful?
When it comes to small DNA segments, we’ve heard the “glass half empty” version of the story many times. Here’s the other side of that story.
Submitted for your consideration: A pair of third cousins twice removed and their 6 cM connection…
According to AncestryDNA, Bryan Smith and his cousin, K, share 6 cM of DNA across 1 segment. And according to Ancestry’s ThruLines, Bryan and Cousin K share a pair of third great grandparents, Reuben Willis Smith and his wife Mary Connell.

The cM value is certainly consistent with the identified relationship but did Bryan and Cousin K inherit their shared DNA from the Smith ancestry as shown? Is the 6 cM segment even valid or could it be an artifact of an imperfect DNA matching algorithm?
Let’s start with an easy evaluation: the shared match list.
Among Bryan and K’s list of shared matches at AncestryDNA:
HG, a descendant of Reuben and Mary’s son, Charles Thomas Smith (HG shares 57 cM with Bryan)
RR, a descendant of Reuben and Mary’s daughter, Fannie Janes Smith (RR shares 47 cM with Bryan and 38 cM with K)
IG, another descendant of Reuben and Mary’s son, Charles Thomas Smith (IG shares 47 cM with Bryan and 71 cM with K)
And at least three other descendants of Reuben and Mary are on the shared match list.
So we’re off to a promising start. In addition to the fact that Bryan and K share DNA and a paper trail leading to Ruben and Mary, this group of matches gives us more evidence suggesting that Bryan and K might be related as suspected.
But what about that 6 cM segment shared by Bryan and K? Is it valid? Did it come from the shared Smith ancestors or did it originate elsewhere?
To get the most comprehensive help in answering these questions, we turn to GEDmatch. As indicated in the ThruLine image above, both Bryan and his father are related to DNA Cousin K through their Smith line. And because K is on GEDmatch, we can see that Bryan and his father both share DNA with K on a specific portion of Chromosome 12:
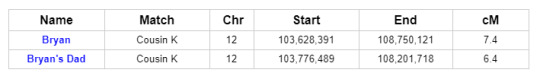
Further investigation reveals that two other descendants of Reuben and Mary, Cousins I and G, share DNA with Bryan and his father on Chromosome 12 in roughly the same location. In fact, all of the matches in question match each other on Chromosome 12:
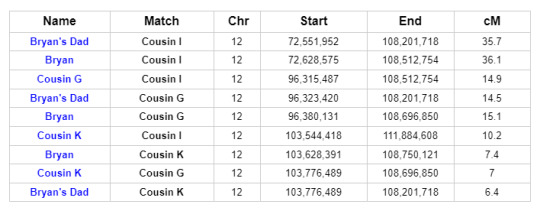
This is what we call a Triangulation Group. It brings the possible genetic connections into sharper focus.
The common segment shared by all of the members of this Triangulation Group indicates that they all share a common ancestor. And we’ve already identified shared ancestry through the Smith line. Cousins I and G are first cousins once removed and they are descendants of Reuben and Mary’s son Charles Thomas Smith...
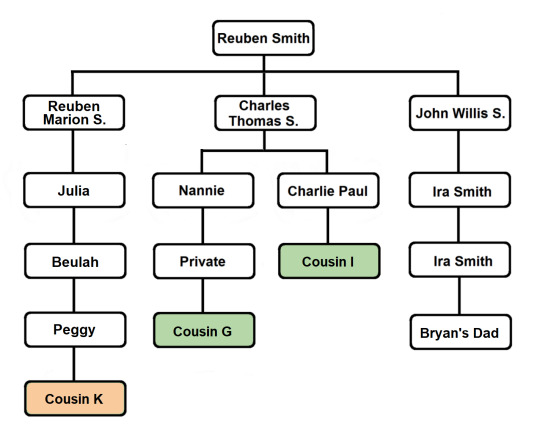
A review of the pedigrees of the matches in question reveals no lines of shared ancestry other than the known shared Smith line. This investigation is summarized briefly in the table below, listing 2nd great grandparent surnames and shared ancestors (blue for paternal names and surnames, light red for maternal names and surnames):
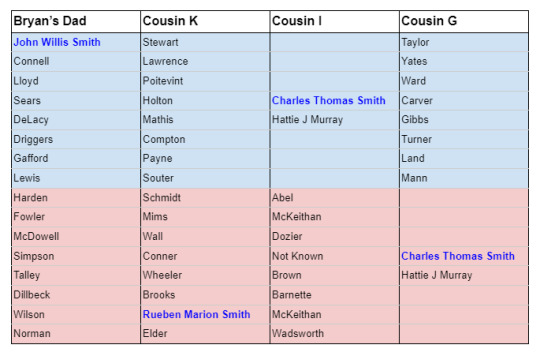
Although we cannot say with perfect certainty that there is no additional common ancestry that conceivably could account for the shared segment of DNA on Chromosome 12, the known evidence doesn’t leave room for much doubt.
For completeness, here’s a chart summarizing the amount of DNA shared by the relatives in question:

And cluster analysis for Cousin G yields a cluster with eight descendants of Reuben Willis Smith, including Bryan Smith and Cousin K:

Not everyone will feel the need to go this far to investigate a 6 cM connection. But this post provides examples of ways to investigate the validity of an ordinary small segment and to determine whether the shared DNA legitimately belongs with the presumed paper trail source of the DNA.
Discussion
Skepticism regarding small segments of shared DNA is appropriate. In comparison to larger shared segments, such segments are more likely to be IBS (false). Additionally, even when small segments can be shown to be reliable, we have to grapple with the fact that small segments can be too old to fall within the reach of reliable historical documentation.
With the exponential growth of the DNA matching databases, the impetus to explore distant matches waned. Reluctance to do the strenuous work involved in using small segments grew. With access to strong genetic connections leading back to target ancestors, why bother with low cM connections?
The sentiment is understandable!
On the other hand, I believe that excessive skepticism has impeded progress in genetic genealogy. As databases have grown, our opportunities for research have multiplied and our research techniques have improved. But at the same time, goalposts for small segment success have been moved to poorly-defined and very unreasonable points.
[From the skeptics: Your success with a small segment doesn’t count if you find a larger segment in a relative! I don’t want to hear about triangulation! Visual phasing is not allowed!]
If we applied such arbitrary restrictions to all areas of genealogy, we’d struggle to get our work done!
Even with our luxuriously large DNA databases, distant genetic connections are the only connections available in some areas of investigation (or to people who hail from less heavily-tested populations). Defeatist refusal to accept low cM matches as evidence in genetic genealogy needlessly limits our potential.
Don’t get me wrong, I’m fully in favor of scholarly rigor. But let’s not allow skepticism to pave the way for denialism!
When distant genetic connections are found to be of dubious quality, they should be set aside. But shared segments should not be judged on the basis of size alone. Even the most fervent opponents of small-segment research will admit that small segments are often valid (IBD). And while these opponents frequently cite IBD/IBS percentages, they ironically fail to see that our ability to find these percentages points directly to a practical method for sorting distant matches on an individual basis.
We are privileged to have access to enormous databases of incredibly valuable genetic information. More than a statistical hiccup that can lead us serendipitously to more reliable information, small DNA segments are messages we carry with us every day, testifying to our connections with our ancestors. Genetic information, even in small amounts, can be just as valuable as any other form of information. We should be good stewards of that information and we should invest good faith effort in understanding how our distant matches can inform us about our rich ancestral history.
I’ll close with this analogy for small segments:
You want some refreshing water but the glass is only half-full. Drink it or toss it out?

Posted with Bryan Smith’s permission. 17 May 2023
#genetic genealogy#DNA testing#DNA#small segments#visual phasing#Triangulation#dna segment#DNA segment triangulation#DNA clustering#clustering#pedigree#family tree#ThruLines
3 notes
·
View notes
Text
When Cavetown says “My brothers are circles, my sisters quadrilaterals” in Taking Care of Things (Lemon Boy album) is that a reference to being transgender and pedigrees or am I just learning about pedigrees in bio and making unnecessary connections
#that line as always confused me idk#I’ve assumed it’s about gender non-conformism forever but I just made that pedigree connection and idk if it’s true but#females are circles and males and squares when representing on pedigrees so it means that to me now#cavetown#cavetown lyrics#lemon boy#taking care of things#pedigree#biology
6 notes
·
View notes

